Written by: Katherine Eaton
An important foundation of my dissertation is the acquisition of metadata from online repositories. I often spend my time working with records of infectious disease outbreaks where the metadata can take the form of:
- Date a pathogen was isolated from the host (ex. outbreak timing)
- Geographic coordinates (ex. spatial spread)
- Disease status (ex. virulence and mortality)
- Download links to genetic sequences (ex. analysis)
The main repository I work with is the National Center for Biotechnology Information (NCBI), one of the most important resources for health informatics. I’d previously written a prototype program to query the repository, fetch records, and organize them into a SQL database. My last blog post showcased a possible application in which I explored ‘secret’ infectious disease datasets that were never published upon and hidden from academic literature. I also undertook some GIS training to be able to plot this information and visualize outbreaks on a global scale. Overall it was a successful endeavor, and has become useful to me as both a time-saving tool in metadata collection, as well as a methodological approach to data discovery.
However, my initial prototype was written so that I, and only I, could use it. In additional, it was not easy to change search queries, or request different metadata fields to be pulled from it. And that was the biggest problem I needed to sort out. So this update is all about the software development that has got me back on track to continue my applications.
Progress #1
I changed the implementation of the program so that it took as input a configuration file, which had two main benefits:
- The command-line usage was much cleaner and simplified (always a good thing)
- The query-and-retrieve algorithm is now flexible and can theoretically retrieve any metadata field the repository provides.
Progress #2
I wrote a module so that a user can annotate the database with their own curated metadata, in the case of missing or incorrect information. This was extremely fun, as it’s a completely dynamic algorithm (note: not dynamic-programming though) and involved teaching the computer to create and name its own variables on the fly. I imagine this opens up security vulnerabilities though, but for a first draft this was a very fun way to get creative in what is otherwise a very boring field of database programming.
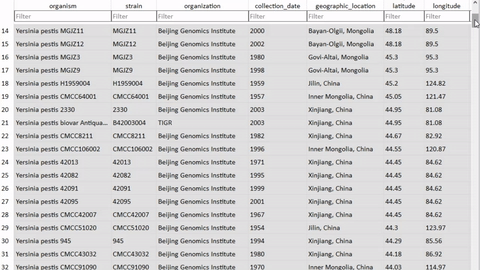
(Yersinia pestis is the name of the bacteria that causes the disease Plague, which is the focus of my dissertation)
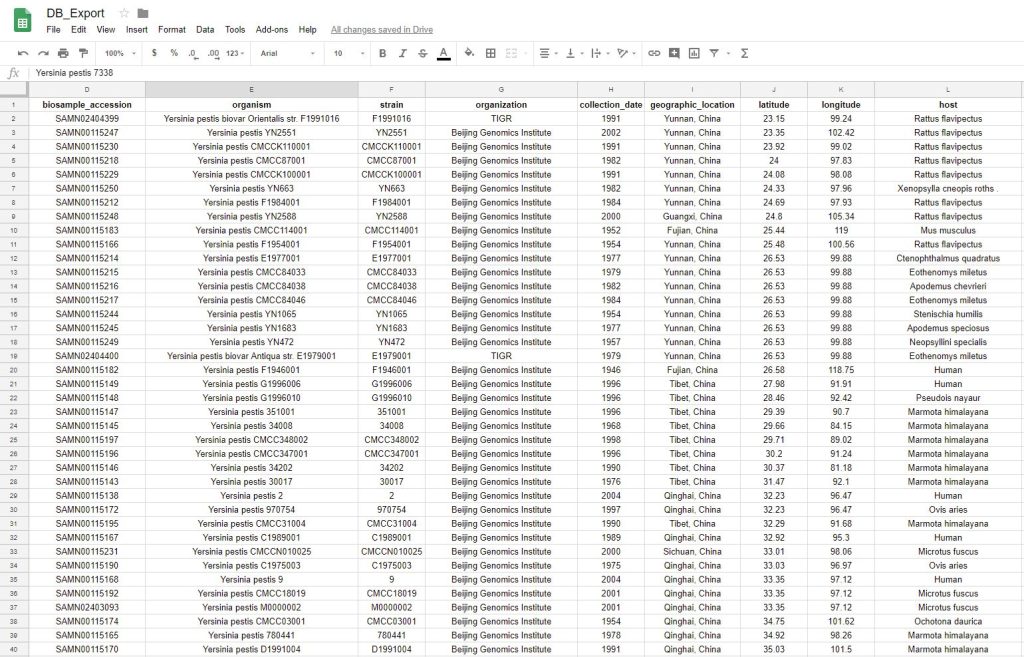
I also wrote a module that would export the SQL database into a plain text file, which could be read by your favorite text editor or manipulated as a spreadsheet. I figured this would be useful for either:
- People who do not want to/do not know how to work with SQL.
- People who want to manipulate the database with downstream text-processing tools.
Progress #3
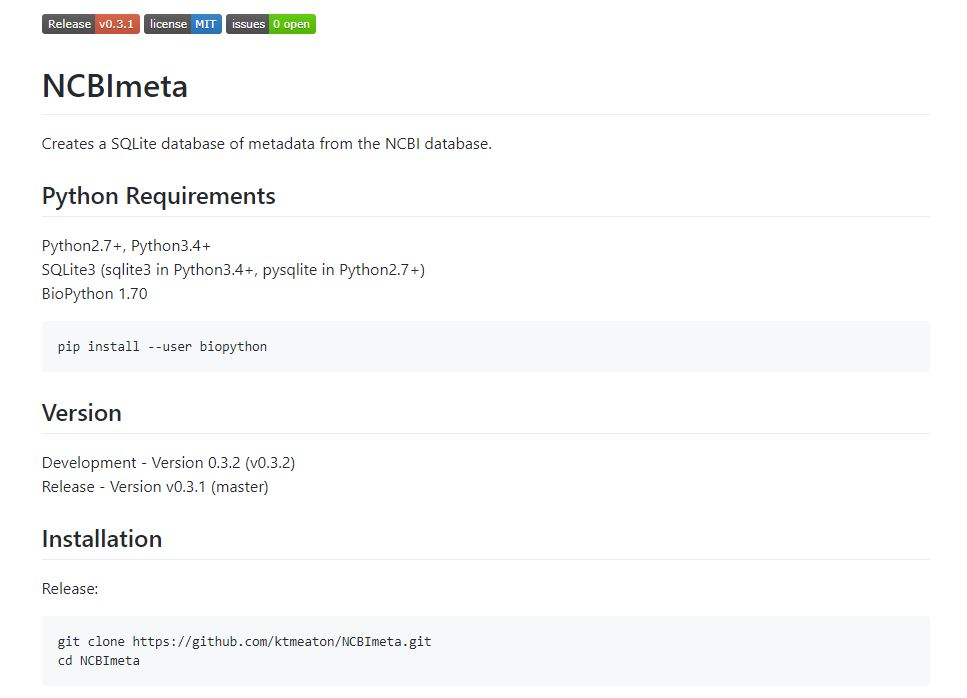
Since I would like to ask people I know to test this run, I improved the documentation for first-time users. I built a fully-functioning landing page on github (https://github.com/ktmeaton/NCBImeta) to introduce new users to the program which will eventually include a full manual and tutorial .
Moving Forward
There are 3 objectives I want to focus on moving forward:
- Improve documentation and clean-up messy code.
- Ask some colleagues to help in testing and finding bugs.
- Applications!
Now that the database can be created with greater flexibility and ease (and with improved retrieval of latitude/longitude coordinates), I will be returning to GIS-based analysis for the next several months. I’ve newly discovered an excellent spatial analysis tool that can help reconstruct how infectious diseases may be evolving and mutating across landscapes (GenGIS). So next time we’ll be back to map construction and testing spatial hypotheses!


Leave a Reply